Recent advances in Nanotechnology allow sophisticated synthesis of nanomaterials with well-defined composition, aspect ratio, size, surface chemistry and functionalization. Biomedical applications of engineered nanomaterials or nanomedine include imaging, sensing, targeting, drug and gene delivery, and therapeutics in diagnosis, prognosis, and treatment of human diseases. Upon entering biological media such as the bloodstream, a nanoparticle forms molecular complexes with encountered proteins, termed as the protein corona. The protein-nanoparticle complex instead of the original nanoparticle dictates its biological identity and function. In addition, proteins in the corona can undergo conformational changes, resulting into either lost or gain of functions. Hence, understanding the structure and dynamics of a nanoparticle–protein corona is essential for predicting the fate, transport, and toxicity of nanomaterials in living systems and for enabling the vast applications of nanomedicine.
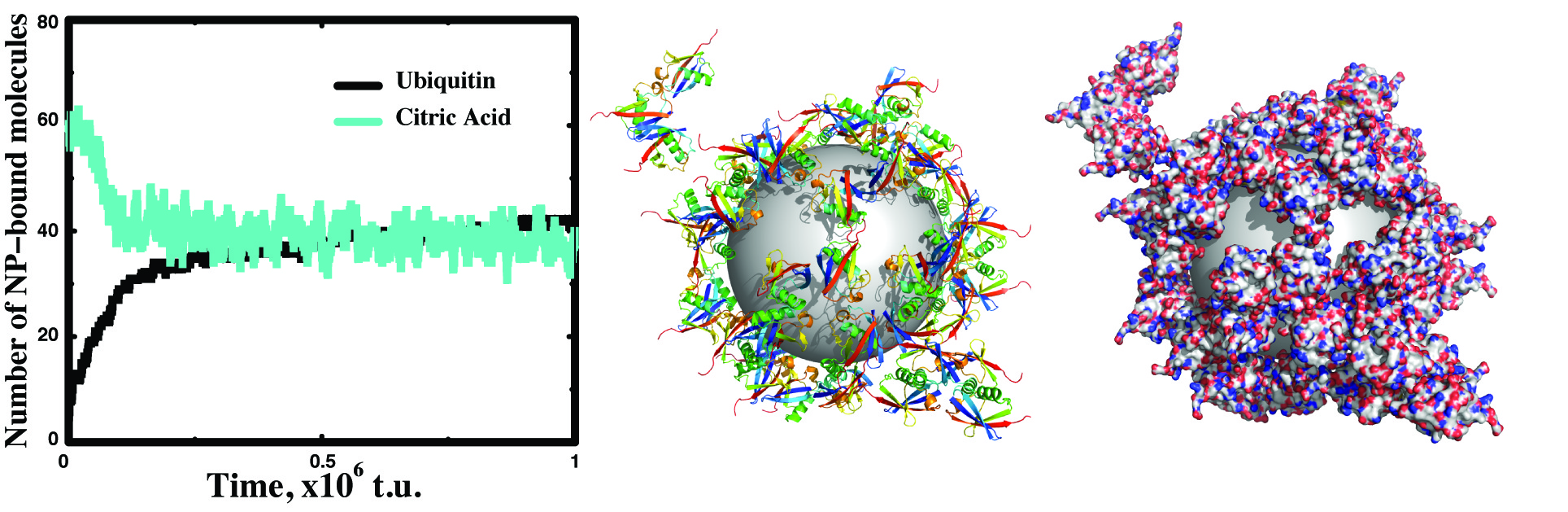
Our group is the first to demonstrate the formation of protein-nanoparticle corona at both atomic and molecule levels in silico using multiscale discrete molecular dynamics (DMD) simulations. Computational results of the ubiquitin-silver nanoparticle (AgNP) corona agreed with high-resolution NMR experiments of the binding between ubiquitin and AgNP, the stretched exponential binding kinetics, and the secondary structure changes of ubiquitin in the corona. Over the years, we have been expanding the capability of DMD to model various types of nanomaterials ranging from carbon-based and polymeric to metal and metal oxide nanoparticles. Using DMD simulations, we have been studying the fundamental interactions between nanoparticles and biomolecules, and trying to uncover the physicochemical determinants of nanoparticles on the self-assembly of biomolecules in the corona.
49. X. Liang, N. Andrikopoulos, H. Tang, Y. Wang, F. Ding* and P.C. Ke*, “Nanoplastic stimulates the amyloidogenesis of Parkinson’s alpha-synuclein NACore”, Small, in press (2023) doi:10.1002/smll.202308753
48. Andrikopoulos N., Li Y., Nandakumar A., Quinn J., Davis T., Ding F.*, Saikia N.*, Ke P.C.*, “Zinc-Epigallocatechin-3-gallate Network-Coated Nanocomposites against the Pathogenesis of Amyloid-Beta”, ACS Applied Materials & Interfaces, 15, 6, 7777–7792 (2023) doi:10.1021/acsami.2c20334
47. J. He, L. Zhou, G. Huang, J. Shen, W. Chen, C. Wang, A. Kim, Z. Zhang, W. Cheng, S. Dai, F. Ding,* and P. Chen*, “Enhanced Label-Free Nanoplasmonic Cytokine Detection in SARS-CoV-2 Induced Inflammation Using Rationally Designed Peptide Aptamer”, ACS Applied Materials & Interfaces, 14, 43, 48464–48475 doi:10.1021/acsami.2c14748 (2022)
46. W. Wei, Y. Li, M. Lee, N. Andrikopoulos, S. Lin, C. Chen, D. Leong, F. Ding, Y. Song, and P. Ke, “Anionic Nanoplastic Exposure Induces Endothelial Leakiness”, Nature Communications, 13: 4757, doi:10.1038/s41467-022-32532-5 (2022)
45. J. Ren, K. Velonia, N. Andrikopoulos, H. Tang, F. Ding, P. C. Ke, C. Chen, “Chemical and Biophysical Signatures of the Protein Corona in Nanomedicine”, Journal of The American Chemical Society, 144(21): 9184–9205 (2022) doi:10.1021/jacs.2c02277
44. H. Tang, Y. Li, A. Kakinen, N. Andrikopoulos, Y. Sun, E. Kwak, T. P. Davis, F. Ding* and P. C. Ke*, “Graphene Quantum Dots Obstruct the Membrane Axis of Alzheimer’s Amyloid Beta”, Physical Chemistry Chemical Physics, 24, 86-97 (2022) doi: 10.1039/D1CP04246G
43. A. Nandakumar, W. Wei , G. Siddiqui, H. Tang, Y. Li , A. Kakinen, X. Wan, K. Koppel, S. Lin, T. Davis, D. Leong, D. Creek, F. Ding, Y. Song, P. C. Ke, “Dynamic Protein Corona of Gold Nanoparticles with An Evolving Morphology”, ACS Appl. Mater. Interfaces, 13, 48, 58238–58251 (2021) doi: 10.1021/acsami.1c19824
42. M. Lee,† N. Ni,† H. Tang,† Y. Li,† W. Wei, A. Kakinen, X. Wan, T. P. Davis, Y. Song,* D. T. Leong,* F. Ding* and P. C. Ke*, “A Framework of Paracellular Transport via Nanoparticles-Induced Endothelial Leakiness”, Advanced Science, 2102519 (2021) doi:10.1002/advs.202102519
41. H. Tang, Y. Li, A. Kakinen, N. Andrikopoulos, Y. Sun, E. Kwak, T. P. Davis, F. Ding* and P. C. Ke*, “Graphene Quantum Dots Obstruct the Membrane Axis of Alzheimer’s Amyloid Beta”, Physical Chemistry Chemical Physics, in press (2021) doi: 10.1039/D1CP04246G
40. A. Nandakumar, W. Wei , G. Siddiqui, H. Tang, Y. Li , A. Kakinen, X. Wan, K. Koppel, S. Lin, T. Davis, D. Leong, D. Creek, F. Ding, Y. Song, P. C. Ke, “Dynamic Protein Corona of Gold Nanoparticles with An Evolving Morphology”, ACS Appl. Mater. Interfaces, in press (2021) doi: 10.1021/acsami.1c19824
39. M. Lee,† N. Ni,† H. Tang,† Y. Li,† W. Wei, A. Kakinen, X. Wan, T. P. Davis, Y. Song,* D. T. Leong,* F. Ding* and P. C. Ke*, “A Framework of Paracellular Transport via Nanoparticles-Induced Endothelial Leakiness”, Advanced Science, 2102519 (2021) doi:10.1002/advs.202102519
38. N. Andrikopoulos, Z. Song, X. Wan, A. Douek, I. Javed, C. Fu, X. Changkui, Y. Xing, F. Xin, Y. Li, A. Kakinen, K. Koppel, R. Qiao, A. Whittaker, J. Kaslin, T. Davis*, Y. Song*, F. Ding*, P.C. Ke*, “Inhibition of Amyloid Aggregation and Toxicity with Janus Iron Oxide Nanoparticles”, Chem. Mater., 33, 16, 6484–6500 (2021) doi: 10.1021/acs.chemmater.1c01947
37. Y. Li, H. Tang, H. Zhu, A. Kakinen, D. Wang, N. Andrikopoulos, Y. Sun, A. Nandakumar, E. Kwak, T. Davis, D. Leong, F. Ding, P. C. Ke, “Ultrasmall Molybdenum Disulfide Quantum Dots Cage Alzheimer’s Amyloid Beta to Restore Membrane Fluidity”, ACS Appl. Mater. Interfaces, 13(25): 29936–29948 (2021) doi: 10.1021/acsami.1c06478
36. Chen P, Ding F, Cai R, Javed I, Yang W, Zhang Z, Li Y, Davis TP, Ke PC, Chen C., “Amyloidosis Inhibition, a New Frontier of the Protein Corona”, Nano Today, 35:100937 (2020) doi: 10.1016/j.nantod.2020.100937
35. Koppel K, Tang H, Javed I, Parsa M, Mortimer M, Davis TP, Lin S, Chaffee AL, Ding F and Ke P, “Elevated Amyloidoses of Human IAPP and Amyloid Beta by Lipopolysaccharide and Their Mitigation by Carbon Quantum Dots”, Nanoscale, in press (2020)
34. Y. Xing, Y. Sun, B. Wang, & F. Ding, “Morphological Determinants of Carbon Nanomaterial-Induced Amyloid Peptide Self-Assembly”, Frontiers in Chemistry, in press (2020)
33. Z Huma,I Javed, Z Zhang, H Bilal, Y Sun, SZ Hussain, TP Davis, DE Otzen, CB Landersdorfer, F Ding, I Hussain and PC Ke, “Nano Silver Mitigates Biofilm Formation via FapC Amyloidosis Inhibition”, Small, 1906674-1906683 (2019) DOI: 10.1002/smll.201906674
32. A Faridi, Y Sun, M Mortimer, RR Aranha, A Nandakumar, Y Li, I Javed, A Kakinen, Q Fan, AW Purcell, TP Davis,* F Ding,* P Faridi,* and P Ke*, “Graphene quantum dots rescue protein dysregulation of pancreatic β-cells exposed to human islet amyloid polypeptide”, Nano Research, in press (2019)
31. I Javed, G Peng, Y Xing, T Yu, M Zhao, A Kakinen, A Faridi, CL Parish, F Ding*, TP Davis*, P Ke* and S Lin*, “Inhibition of Amyloid Beta Toxicity in Zebrafish with A Chaperone-Gold Nanoparticle Dual Strategy”, Nature Commun, in press (2019)
30. Y. Sun, A. Kakinen, C. Zhang, Y. Yang, A. Faridi, T. P. Davis, W. Cao, P. C. Ke and F. Ding, “Amphiphilic Surface Chemistry of Fullerenols Is Necessary for Inhibiting the Amyloid Aggregation of Alpha-Synuclein NACore”, Nanoscale, in press, (2019)
29. P. C. Ke, E. H. Pilkington, Y. Sun, I. Javed, A. Kakinen, G. Peng, F. Ding, . P. Davisa, “Mitigation of Amyloidosis with Nanomaterials”, Advanced Materials, in press, (2019)
28. M. Wang, Y. Sun, X. Cao, G. Peng, I. Javed, A. Kakinen, T.P. Davis, S. Lin, J. Liu, F. Ding, and P. Ke, “Graphene Quantum Dots against Human IAPP Aggregation and Toxicity in Vivo”, Nanoscale 10, 19995, DOI: 10.1039/C8NR07180B (2018)
27. A. Faridi,Y. Sun, Y. Okazaki, G. Peng, J. Gao, A. Kakinen, P. Faridi, M. Zhao, I. Javed, A.W. Purcell, T.P. Davis, S. Lin, R. Oda, F. Ding, P. Ke, “Mitigating Human IAPP Amyloidogenesis in Vivo with Chiral Silica Nanoribbons”, Small, 14, 1802825, DOI: 10.1002/smll.201802825 (2018)
26. B. Wang, Y. Sun, T. Davis, P.C. Ke, Y. Wu, and F. Ding, “Understanding the Effects of PAMAM Dendrimer Size and Surface Chemistry on Serum Protein Binding with Discrete Molecular Dynamics Simulations”, ACS Sustainable Chemistry & Engineering, 6 (9),11704–11715, DOI: 10.1021/acssuschemeng.8b01959 , (2018)
25. Pilkington E.P., Gustafsson O.J.R., Xing Y., Hernandez-Fernaud J., Zampronio C., Kakinen A., Faridi A., Ding F., Wilson P., Ke P.C. and Davis T.P., “Profiling the Serum Protein Corona of Fibrillar Human Islet Amyloid Polypeptide”, ACS Nano, 12, 6066−6078 (2018) DOI: 10.1021/acsnano.8b02346
-Featured as Journal Front Cover .
24. A. Kakinen, J. Adamcik, B. Wang, X. Ge, R. Mezzenga, T.P. Davis, F. Ding, and P. Ke, “Nanoscale inhibition of polymorphic and ambidextrous IAPP amyloid aggregation with small molecules”, Nano Research, 11(7): 3636–3647 (2017)[download]DOI: 10.1007/s12274-017-1930-7
23. I. Javed, Y. Sun, J. Adamcik, B. Wang, A. Kakinen, E. Pilkington, F. Ding, R. Mezzenga, T. Davis, Thomas; P. Ke, “Co-fibrillization of pathogenic and functional amyloid proteins with gold nanoparticles against amyloidogenesis”, Biomacromolecules, in press (2017) DOI: 10.1021/acs.biomac.7b01359
22. E. Pilkington, M. Lai, X. Ge, W. Stanley, B. Wang, M. Wang, A. Kakinen, M. Sani, M. Whittaker, E. Gurzov, F. Ding, J. Quinn, T. Davis, P. Ke, “Star Polymers Reduce IAPP Toxicity via Accelerated Amyloid Aggregation”, Biomacromolecules, in press (2017)
21. J. Yang, B. Wang, Y. You, W. Chang, K. Tang, Y. Wang, W. Zhang, F. Ding, S. Gunasekaran, “Probing the modulated formation of gold nanoparticle-beta lactoglobulin corona complexes and its applications”, Nanoscale, in press (2017) [link]
20. B. Wang, E.H. Pilkington, Y. Sun, T.P. Davis, P.C. Ke and F. Ding, Modulating protein amyloid aggregation with nanomaterials, Environmental Science Nano, in press (2017)
19. N.K. Geitner, W. Zhao, F. Ding, W. Chen, M. R. Wiesner, “Mechanistic Insights from Discrete Molecular Dynamics Simulations of PesticideNanoparticle Interactions”, Environmental Science & Technology, in press (2017) [link]
18. S. Lin, M. Mortimer, R. Chen, A. Kakinen, J.E. Riviere, T.P. Davis, F. Ding,* and P. C. Ke*, “NanoEHS beyond Toxicity – Focusing on Biocorona”, Env. Sci. Nano, in press (2017) [link]
17. Pilkington E.H., Xing Y., Wang B., Kakinen A., Wang M., Davis T.P., Ding F., Ke P.C., “Effects of Protein Corona on IAPP Amyloid Aggregation, Fibril Remodelling, and Cytotoxicity”, Scientific Reports, in press (2017)
16. B. Wang,T. Blin, A. Käkinen, X. Ge, E.H. Pilkington, J.F. Quinn, M.R. Whittaker, T.P. Davis, P.C. Ke and F. Ding, “Brushed polyethylene glycol and phosphorylcholine for grafting nanoparticles against protein binding”, Polymer Chemistry, in press (2016)
15. T. Blin, A. Kakinen, E.H. Pilkington, A. Ivask, F. Ding, J.F. Quinn. M.R. Whittaker, P.C. Ke and T.P. Davis, “Synthesis and In Vitro Properties of Iron Oxide Nanoparticles Grafted with Brushed Phosphorylcholine and Polyethylene Glycol”, Polymer Chemistry, in press (2016)
14. E.N. Gurzov, B. Wang, E.H. Pilkington, P. Chen,A. Kakinen, W.J. Stanley, S.A. Litwak, E.G. Hanssen,T.P. Davis, F. Ding, and P.Chun Ke, “Inhibition of hIAPP Amyloid Aggregation and Pancreatic β-cell Toxicity by OH-terminated PAMAM Dendrimer”, Small, in press (2016)
13. S. Radic, T.P. Davis, P.C. Ke and F. Ding, “Contrasting effects of nanoparticle-protein attraction on amyloid aggregation”, RSC Advances, in press (2015)
12. P. Nedumpully-Govindan, E.N. Gurzov, P. Chen, E.H. Pilkington, W.J. Stanley, S.A. Litwak, T.P. Davis, P.C. Ke, and F. Ding, “Graphene Oxide Inhibits hIAPP Amyloid Fibrillation and Toxicity in Insulin-Producing NIT-1 Cells”, PCCP, in press (2015)
11. Wang B, Geitner NK, Davis TP, Ke PC, Ladner DL and Ding F, “Deviation from the Unimolecular Micelle Paradigm of PAMAM Dendrimers Induced by Strong Inter-Ligand Interactions”, Journal of Physical Chemistry C, in press (2015)
10. Ge XW, Ke PC, Davis TP and Ding F, “A Thermodynamics Model for the Emergence of a Stripe-like Binary SAM on a Nanoparticle Surface”, Small, in press (2015)
9. DeFever R., Geitner N., Bhattacharya P., Ding F., Ke P.C., Sarupria S., “PAMAM dendrimers and graphene: Materials for removing aromatic contaminants from water”, Environmental Science & Technology, in press (2015)
8. Geitner N., Wang B., Andorfer R.; Ladner D.; Ke P.K.; Ding F., “The structure-function relationship of PAMAM dendrimers as robust oil dispersants”, Environmental Science & Technology, 48(21):12868-75, (2014)
7. Wang B., Seabrook S.A., Nedumpully-Govindan P., Chen P., Yin H., Waddington L., Epa V.C., Winkler D.A., Kirby J.K., Ding F., Ke P.C., “Thermostability and Reversibility of Silver Nanoparticle-Protein Binding”, Physical Chemistry Chemical Physics, in press (2014)
6. S. Radic, P. Nedumpully-Govindan,R. Chen, E. Salonen, J.M. Brown, P.C. Ke, and F. Ding, “Effect of Fullerenol Surface Chemistry on Nanoparticle Binding-induced Protein Misfolding”, Nanoscale, in press (2014)
5. Y. Wen, N.K. Geitner, R. Chen, F. Ding, P. Chen, R.E. Andorfer, P.N. Govindan, and P.C. Ke, Binding of Cytoskeletal Proteins with Silver Nanoparticles, RSC Adcances, in press, (2013).
4. EE. Salonen, F. Ding, and P.C. Ke, “Fate, behavior and biophysical modeling of nanoparticles in living systems” in “Engineered Nanoparticles and the Environment: Biophysicochemical Processes and Toxicity”, Edited by B. Xing, C.D. Vecitis, & N. Senesi, Wiley-IUPAC (2016) [download]
3. A. Kakinen, F. Ding, P. Chen, A. Kahru*, and P.C. Ke, “Interaction of Silver Nanoparticles and Firefly Luciferase and Its Impact on Enzyme Luminescence”, Nanotechnology, in press. (2013)
2. S. Radic, N. Geitner, R. Podila, A. Kakinen, P. Chen, P.C. Ke*, and F. Ding*, “Competitive Binding of Natural Amphiphiles with Graphene Derivatives”, Scientific Reports, in press. (2013)
1. F. Ding*, S. Radic, R. Chen, P. Chen, J.M. Brown and P.C. Ke*, “Direct observation of silver nanoparticle-ubiquitin corona formation”, Nanoscale, in press (2012)
